To recognize genes accountable for that similarities of NB and PCC gene expression patterns, we now have searched for genes expressed in both tumors with all the lowest fold modify and variation. The previously published reference genes were filtered out from these gene lists to pick genes with unchanged expres sion exclusively in between NB and PCC. The examination was carried out on three different platforms. By this strategy we have now identified 62 genes that had been popular in at least two comparisons. By IPA of these gene sets, death receptor signaling was quite possibly the most substantial popular pathway in every comparison. Distinctions in between NB and PCC tissues The comparison of NB and PCC was doable on 3 platforms, and through the comparison of these gene lists we have now identified 758 popular expression changes in a minimum of two comparisons, for instance, anaplastic lymphoma receptor tyrosine kinase, neurotrophic tyrosine kinase, receptor, sort three, tubulin, beta 2A C, 3.
By IPA, cancer regulation by stathmin1 selleck chemicals has been identified because the most sizeable pathway involving PCC and NB. Expression modifications among distinctive subgroups of NB The comparison of significant gene lists among phases 1 and 4 was probable on 3 distinctive platforms. Within these, we now have uncovered 28 common expression modifications in not less than two scientific studies. The comparison of stage 4S and stage 4 NB was probable on one particular platform. The record of considerably differentially expressed genes was when compared to the gene checklist reported by Schramm et al. and 5 common genes are actually discovered in both, such as, aldehyde dehydrogenase 3 family member A2 and phosphoinositide 3 kinase regulatory subunit 1.
The examination selleck chemicals DZNeP of gene expression profiles of MYCN non amplifying and MYCN amplifying NB was doable on three platforms. The comparison of these uncovered 259 typical important expression modifications in no less than two studies. To identify the most trusted expression improvements, we intersected this gene set having a further a single, reported by Schramm et al. By this method, we’ve got recognized 9 genes with major expression changes that were widespread in all 4 gene sets, for ex ample, DEAD box polypeptide one, MYCN, orni thine decarboxylase 1, transketolase. Through the comparison of gene lists exposed from your stat istical analysis of early versus late stage, MYCN non amplifying versus MYCN amplifying and favorable versus unfavorable NB samples and in the literature search, we now have uncovered 519 genes that have been reported at the very least twice in these gene lists. Amongst these, probably the most prevalent genes were, NTRK1, MYCN, secretogranin II, pleiotrophin, neuronal cell adhesion molecule and ODC1. 
Monthly Archives: May 2014
7 1 1 and also the decarboxylation of pyruvate to acetaldehyde
seven. 1. 1 as well as the decarboxylation of pyruvate to acetaldehyde. Moreover, nearly all of the increase in tolerance for the WOAs may very well be attributed to these two reactions, in contrast to the effects underneath histidine starvation the place the response was distributed among various reactions. Notably, the overall gene expression changes of these reactions had been of fairly modest magni tude in half of your scenarios. To put this in perspec tive, when the reactions with gene associations had been sorted by descend ing buy of magnitude of their all round gene expression changes, reaction EC 4. 1. one. one ranked beneath 10 in three out of the four cases. Note that we didn’t include things like during the evaluation the gene expression improvements linked to the efflux processes of protons and carboxylic anions.
Moreover the two most influential reactions, only a few many others had a significant personal contribution for the predicted tolerance. This outcome is in agreement with the sloppiness residence, inside the sense that a networks function is established by a decreased quantity of parameters. Sensitivity and robustness of model Vismodegib price predictions An advantage of our method is it’s a small variety of standard fitting parameters, B, and. We investigated the effect of those fitting parameters on model predictions by simulating the model for different values of those param eters. For that WOA therapy experiments, we determined the normalized SSE between the predicted and experimen tal values of your exchange fluxes and biomass yield as being a function of every parameter. The analysis showed the predicted fluxes have been robust with respect to alterations in these parameters.
The simulations of your histidine starvation experiments have been fairly robust to variations MK2206 in these parameters all around the values reported in Table one, as proven in Additional File one. We also investigated in the event the system was sensitive for the input gene expression information or if your benefits might be EC obtained with arbitrary data. Briefly, we simulated the designs with data produced by randomly shuffling the original expression data. The simulation results showed that it can be unlikely that very similar success might be obtained making use of random gene expression data. Furthermore, uncertainty propagation examination showed that the process is robust with respect to experimental noise while in the gene expression data and flux distributions. Discussion Lengthy standing barriers impeding the development of big scale kinetic models of metabolic process are staying over include the assist of developments in substantial throughput technologies and computational analyses. Modelers are now faced using the challenge of integrating  the increas ingly out there setting up blocks to make coherent mathem atical representations of biological systems.
the increas ingly out there setting up blocks to make coherent mathem atical representations of biological systems.
The plot additional indicated a detrimental correlation in betw
The plot more indicated a negative correlation between these parameters and the pore water ammonia concentration. The significantly reduce ammonia concentration mea sured from the Troll samples compared on the Oslofjord samples may very well be a end result of your nitrifiers efficient me tabolism of ammonium. Particularly Nitrosopumilus, strain SCM1, is proven to get a higher affinity for ammonia. Interestingly, the PCA plot indicated a powerful constructive correlation concerning Thaumarchaeota as well as the geochemical parameters zinc and calcium. The correlation amongst calcium and Thaumarchaeota could in aspect be explained by the calcium carbonate mound uncovered near to Tpm1 2, in which the Thaumarch aeota had been most abundant. Large variance detected within the Troll area The high variance existing amongst the Troll samples indi cates environmental differences associated for the distinct structures around the seabed from the region.
Interestingly the Tpm1 one and Tpm1 two samples had been dissimilar, probably as a result of pockmarks massive size and heterogeneity. Near to the eastern slope, wherever sample Tpm1 2 was taken, biogenic carbonate structures almost certainly formed throughout former methane seepage may be viewed. Meanwhile, no such automobile bonate structures were detected at selleck chemical the western slope in which sample Tpm1 1 was taken. The PCA analysis positioned Tplain and Tpm1 two consid erably more left along PC1 than the other Troll sam ples. By far the most striking big difference in geochemical composition between Tplain and Tpm1 2 on one particular side and Tpm1 1, Tpm2 and Tpm3 on the other was the considerably reduce concentration of aliphatic hydrocarbons in Tplain and Tpm1 two in contrast to the other Troll samples. This trend was also seen inside the PCA plot. In combination which has a higher taxonomic and meta bolic prospective for hydrocarbon degradation, this indi cates a extra active hydrocarbonoclastic subcommunity in Tplain and Tpm1 2.
Whilst subsystems concerned in degradation of aromatic hydrocarbons have been detected in all metagenomes, major buy inhibitor” overrepresentation in contrast to your Oslofjord metagenomes could only be detected in Tplain and Tpm1 two, thereby supporting a far more active hydrocarbon degrading neighborhood in these samples. The Tplain and especially the Tpm1 two metagenomes also had a larger fraction of reads assigned to critical genes for hydrocarbon degrad ation compared to the other samples. Further, recognized hydrocarbon degrading genera from both Alphaproteobacteria and Gammaproteobacteria have been overrepresented in Tplain and Tpm1 two in contrast to the Oslofjord meta genomes. This trend also can be witnessed during the PCA plot where the parameters Proteobacteria and Metabolic process of Aromatic Compounds are significant con tributors in separating Tplain and Tpm1 two in the other samples.
The protein classification was carried out using the PANTHER Gene
The protein classification was performed making use of the PANTHER Gene and Protein Classification System 62. On line database searches and comparisons for SSDCL 1 and SSHSP90 had been carried out with Integrated Protein Classification database doc. The GenBank accession numbers for the numerous sequence alignment for SSDCL one homologues were, Chae tomium globosum, XP 001223948. 1,Podospora anserina, XP 00190115. 1,N. crassa, XP 961898, Magnaporthea gri sea, A4RKC3. two, Cryphonectria parasitica, Q2VF19. 1, Scler otinia sclerotiorum, XP 001585179. one and Gibberella zeae, XP 389201. one. The GenBank accession numbers for your numerous sequence alignment for SSHSP90 homologues had been, P. brasiliensis, AAX33296. 1, P. anserina, XP 00 19911127. 1, A. nidulans, XP 681538. one, Ajellomyces der matitidis, XP 002624667. one, Phaeosphaeria nodorum, XP 001791544. 1, S. cerevisiae, EGA76545. 1, Grosmannia clavigera, EFX01463.
1, Homo sapiens, NP 005339. Background The Coal Oil Point seep spot, found in the Santa Barbara Channel, California, selelck kinase inhibitor is amongst the most lively seep places in the globe. Seepage on the green residence gasoline methane as well as other hydrocarbons has occurred within this location for in excess of 500 000 many years. The methane emitted through the COP is largely of thermogenic origin as well as daily emission continues to be esti mated to become not less than forty metric tons. At a global scale, the oceans only make up about 2% on the global methane emission price range. This low degree is explained by prokaryotic oxidation of methane in marine sediments and bedrocks ahead of it reaches the water column. The oxygen penetration level in marine sediments is shallow, so the majority of the methane oxidation takes spot at anaerobic problems. Anaerobic oxidation of methane is assumed for being a coupling of reversed metha nogenesis and sulphate reduction.
This approach is probable performed by the but uncultured anaerobic methano trophic archaea in syntrophy with sulphate reducing bacteria. Determined by phylogeny, ANME may be divided into 3 clades, ANME one, ANME two and ANME three. ANME 2 and ANME three are affiliated to the Methanosarcinales, Ganetespib though ANME one is only dis tantly associated with the Methanosarcinales and Methanomi crobiales. The two ANME one and ANME two are linked with sulphur reducing deltaproteobacteria with the Desulfosarcina/Desulfococcus branch. ANME three is primarily associated with SRB strains closely linked to Desulfobulbus. The reversed methanogenesis model for AOM has gained assistance by a metagenomic examine on ANME at Eel River and sequencing of an ANME 1 draft gen ome.
The coverage of SNP indel matched reads was set as not smaller th
The coverage of SNP indel matched reads was set as not smaller than two. If a SNP indel was identified only from just one read through, it had been regarded as for being very likely from a sequen cing error and as a result not thought to be a authentic SNP indel within this research. To test the accuracy of SNP calling, we created a statistical system to model the sequencing error distribution. The model is described briefly under. According on the Illumina Solexa sequencing technology report, the sequencing error charge should be reduce than 2%, and accordingly, a somewhat stringent sequencing error price, 0. 02, was selected. Provided the complete read coverage of the nu cleotide web page and the substitution coverage, the probability of the nucleotide in a specified internet site currently being brought about by sequencing errors, p, might be simulated like a Poisson distribution, using the single parameter, A nucleotide which has a probability lower compared to the pre defined sizeable level needs to be regarded as being a likely SNP in lieu of a sequencing error.
The p values of likely SNPs have been more corrected with False Discovery Price for a number of statistical tests. Only individuals with corrected p values reduce than 0. 05 had been viewed as to get true SNPs. More than 95% the SNPs detected with all the above described simplified SAMtools based mostly process showed q values reduced than 0. 05. Digital gene expression data processing, virtual tag extraction, the original source and mapping the DGE sequence tags The adapter sequences have been lower through the raw reads using FASTX Toolkit, The remaining tags have been 17 18 nucleotides prolonged. Each and every tag was additional counted by a customized perl script.
Virtual tags from the annotated banana transcriptome, novel transcripts identified from our very own RNA seq final results, plus the Musa genome sequence had been extracted from each up and down Regorafenib structure stream sequences of all NlaIII restriction sites. The downstream tags have been right lower and marked as the sense strand, though the reverse complementary up stream tags had been minimize and marked as antisense strand. The predicted tags were named as cds. tag, novel. tag, and genome. tag, respectively, according towards the refer ence sequences described over. The processed one of a kind sequence tags were mapped to cds. tag initial by BLAST with all the word length 17. The unmapped tags have been gathered and fur ther mapped for the full Musa cds se quences. The remaining unmapped tags had been mapped to novel. tag, the novel transcripts, genome.
tag,  and total genome sequences sequentially. Statistical analysis The Bioconductor package deal DESeq was employed to normalize tag counts and acquire variance stabilized ex pression values for every gene. Pearson correlation coeffi cients were calculated to examine the gene expression data across all the samples using R, We made use of heatmap. two perform in the gplots pack age in R to construct heatmaps of correlation coefficients for all 9 samples, To do away with background noise, the transcript abun dance was set to twenty should the normalized value was below twenty when calculating fold modify for comparison.
and total genome sequences sequentially. Statistical analysis The Bioconductor package deal DESeq was employed to normalize tag counts and acquire variance stabilized ex pression values for every gene. Pearson correlation coeffi cients were calculated to examine the gene expression data across all the samples using R, We made use of heatmap. two perform in the gplots pack age in R to construct heatmaps of correlation coefficients for all 9 samples, To do away with background noise, the transcript abun dance was set to twenty should the normalized value was below twenty when calculating fold modify for comparison.
SP01, the most abundant SP transcript, corresponds to a protein t
SP01, quite possibly the most abundant SP transcript, corresponds to a protein that appears inside the literature below the names of habutobin and flavoxobin, a weakly throm bin like enzyme of 242 amino acids that exclusively releases fibrinopeptide A from fibrinogen, No information and facts is available with regard to attainable kallikrein like activity. On the other hand, Yamamoto et al. found that flavoxobin is definitely an lively C3 convertase that selectively releases C3b and C3a. It remains active in blood containing endogenous protease inhibitors, and promotes massive C3 consump tion, and also to a lesser extent, C5 cleavage. A kinin releasing enzyme, flavoviridiobin, is additionally acknowledged from this venom, even so, due to the fact no sequence data are available, we can not recognize it amid our transcripts.
Enzymatic digests of crude venom effected with trypsin, chymotrypsin, and Glu C yielded peptides that accounted for 94. 6% from the major construction of SP01, Realistic peptide coverage of transcripts as minor as 0. 24% was attained, In contrast for the Protobothrops library, the Ovophis library contained transcripts for 26 different SPs, Peptide coverage of 36% or over was selelck kinase inhibitor attained for 22 of those, with coverage over 70% for eleven of them. Two transcripts seem to get plasminogen activators, while SP20 is most just like a kinin releasing enzyme from your venom of Bothrops jararaca. Serine proteases display many amino acid substitutions, plus the structural determinants that specific ally account for kinin releasing action are unknown, The trouble in assigning pharmacological actions to distinct sequence variations is right away apparent on a cursory examination of More file eleven.
Figure S4 and Added file twelve. Figure S5. Wu et al. reported a novel class of inactive serine protease homologs that displayed an arginine substitution for His 43 with the catalytic triad. SP13 was the DCC-2036 only serine protease in our Protobothrops library that showed this His Arg mutation, nevertheless, the Ovophis library contained 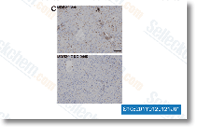 eight transcripts with His X substitutions, Two of those, SP08 and SP22 showed His Lys substitutions. two putative thrombin like enzymes, SP16 and SP17 displayed His Asn substitutions, and SP07 had a His Ala sub stitution. Numerous other sequence differences appear in that transcript too, SPHs from other sources have already been proven to possess diverse activities, so it really is attainable that inactive SPs in venoms have produced other unknown functions, some of which could be specialized for unique prey types. An inactive catalytic triad is but among several structural differences manifested by Ovophis SPHs, Just about every one of the cysteine residues are in vary ent positions likewise, although inside of the group, most residues are conserved across most sequences.
eight transcripts with His X substitutions, Two of those, SP08 and SP22 showed His Lys substitutions. two putative thrombin like enzymes, SP16 and SP17 displayed His Asn substitutions, and SP07 had a His Ala sub stitution. Numerous other sequence differences appear in that transcript too, SPHs from other sources have already been proven to possess diverse activities, so it really is attainable that inactive SPs in venoms have produced other unknown functions, some of which could be specialized for unique prey types. An inactive catalytic triad is but among several structural differences manifested by Ovophis SPHs, Just about every one of the cysteine residues are in vary ent positions likewise, although inside of the group, most residues are conserved across most sequences.
These SEGs can help us to uncover genes correlated with calyx abs
These SEGs can help us to seek out genes correlated with calyx abscission approach. Specially a num ber of stage precise and therapy distinct expressed genes are prone to be major genes related with calyx abscission. Functional annotation of differentially expressed genes Gene Ontology is surely an worldwide standardized gene perform classification program that describes prop erties of genes and their solutions in any organism. Within this study, a total of twelve,054 differentially expressed genes that may be categorized into 41 practical groups had been located, The key subcategories have been as follows. 4 subcategories for cellular element, three subcategories for molecular function, and 7 subcategories for biological process, These outcomes indicate that expressed genes working in binding, catalytic activity, metabolic system, and cellular procedure are significant through the calyx abscission course of action.
Only a number of genes had been clus tered regarding synapse, synapse portion, metallo chaperone activity, biological adhesion and immune strategy process and viral reproduction. To further investigate the function of different expressed genes through calyx abscission, drastically enriched Kyoto Encyclopedia selleckchemRGFP109 of Genes and Genomes pathways were identified according to the P values and enrichment issue. Carrying out a BLAST search towards the KEGG database indicated that expressed genes were concerned in 251 pathways, As shown in Further file three, nine KEGG pathways have been observed to get signifi cantly overrepresented in calyx abscission processes.
People genes correlated with calyx abscission kinase inhibitor Epigenetic inhibitor largely invol ved in photosynthesis, plant hormone signal transduction, cell wall modification, transcriptional regulation and carbohydrate metabolic process had been utilised for subsequent ana lysis. These trends had been consistent with all round deve lopmental actions all through abscission processes, On top of that, a few other biological processes which have not previously been reported to be linked with calyx abscission, such as flavonoid biosynthesis and flavone fla vonol biosynthesis, were noticed considerably changed while in calyx abscission processes. These could possibly be novel genes that are related to your calyx abscission practice in Kuerlexiangli fruit. Cluster of calyx abscission relevant genes Impacts on photosynthesis Reduce in photosynthesis may perhaps be an essential contrib uting issue for that abscission of flowers and fruitlets while in the abscission processes, which was confirmed by our existing experiment. Our benefits showed that 230 genes encoding photosynthesis relevant genes were differential expression underneath numerous therapies, Altered expressions were found for quite a few genes concerned in carbon fixation, photosystem I, photosystem II, and photosynthetic electron transport.
Ear de velopment is often divided into four phases the developme
Ear de velopment is often divided into four stages. the development point elongation phase, spikelet differentiation phase, the floret primordium differentiation phase and floret organ differentiation phase. Plant resources were collected as described previously, Briefly, ears had been manually collected with the 4 developmental phases in accordance to your plant options combined with microscopic observation. Every one of the samples were harvested and instantly frozen in liquid nitrogen and stored at 80 C. The total RNA from each and every sample was then isolated employing Trizol in accordance to the manufacturers directions. Modest RNA library preparation and sequencing The total RNAs were pooled for every of four developmen tal stages for Solexa sequencing.
Right after small RNA cloning, the sequencing inhibitor Dinaciclib procedures have been carried out as described previously, In quick, sequencing was carried out as follows. approximate a hundred ug of complete RNA was purified by polyacrylamide gel electrophoresis, to enrich for molecules while in the variety of 18 30nt, and ligated with adapters towards the five and three terminals of your RNA. Then, minor RNA molecules have been made use of as templates for cDNA synthesis. In total, 18 PCR cycles and agarose gels had been employed for amplification and fragments of around 90 nt in cluding the two compact RNA and adaptors, separately. The purified DNA was made use of Solexa sequence examination per formed by the Illumina platform. Digital top quality information have been created in the picture files made from the sequencer. Just after excellent management working with popular pipeline, clean reads were immediately implemented for even further bioinformatics analysis.
Degradome library construction Modest cDNA libraries implementing the sliced ends of poly adeny lated transcripts from maize ears of four developmental stages were constructed according to past reports, By CHIR-99021 employing the Oligotex kit, 200 ug of total RNA was employed for extracting poly RNA, which have been ligated with an RNA adapter consisting of the MmeI recognition web site in its 3 end. Right after ligation, 1st strand cDNA was produced employing oligod and the PCR item was amplified utilizing five PCR cycles. The PCR item was purified and digested with MmeI. The digested PCR prod uct was then ligated to a double stranded DNA oligo nucleotide with degenerate nucleotides with the five or 3 ends. The ligation merchandise was even further gel purified and amplified applying 10 PCR cycles. The final PCR item was purified and sequenced using Illuminas sequencing by synthesis sequencing engineering.
MiRNA microarray assays MiRNA microarray assays of various developmental phases had been performed by LC Sciences, The custom uparaflo microfluidic chip contained 632 exclusive plant miRNAs of release version 18, representing 1,187 miRNAs from 4 plant species, and 26 further unique miR NAs of maize identified by Solexa sequencing, representing 26 novel miRNAs.
ribicola life cycle These host genes could possibly be located t
ribicola daily life cycle. These host genes could be observed to contribute extra to quantitative disorder resistance. On this examine, about two. 5% of your entire transcriptome assembly was recognized as rust responsive genes and 85% of them were functionally annotated in P. monticola defense, but their putative contribution to host resistance to C. ribicola awaits verification by func tional genomics studies or association studies targeted on exploring gene variation in P. monticola populations. RNA seq examination with the WP BR interactions revealed that two kinds of plant R candidates had been up regulated exclusively in resistant geno variety following C. ribicola infection, suggesting a distinct role of these R candidates in Cr2 mediated resistance.
the biosynthesis and signalling pathways of multiple plant supplier CC-292 hormones were coordinately regulated following rust infection, indicating the auxin and ABA mediated signaling pathways are in volved in white pine resistance to rust. three a set of novel TFs had been identified in response to C. ribicola infection, several of them spe cifically responsive during the incompatible WP BR interaction. and four several households of PR proteins, ROS relevant proteins, UPS proteins, and ret rotransposons have been differentially expressed in the tran scriptional degree amongst resistant and vulnerable genotypes following C. ribicola infection. Solutions Plant materials and rust inoculation Inoculation of P. monticola seedlings with C. ribicola was the exact same as described previously, In short, 200 6 month previous seedlings from the open pollinated seed lot 3926 have been in oculated with C.
ribicola in an inoculation chamber developed to facilitate basidiospore shed with day tem peratures of 16 C and evening temperatures of 12 C, recommended reading respectively. Basidiospores had been shed from C. ribicola infected Ribes nigrum leaves that were laid on the metal mesh over the white pine seedlings. Infected leaves had been collected from a Ribes backyard on Vancouver Island where only pathogenic avcr2 isolates had been out there. Ba sidiospore density was monitored by putting glass slides at random beneath the mesh throughout rust inoculation. WWP needles were inoculated at a spore density higher than three,000 spores per square centimeter. Main nee dles were collected from at the least 10 seedlings individu ally for every remedy at 0 and four dpi and stored at 80 C.
Every single seedling was pre recognized as resistant or susceptible genotype employing Cr2 linked DNA markers as reported previously, cDNA library building Complete RNA was extracted from somewhere around 5 grams of needles pooled from at the very least ten seedlings per treat ment making use of a protocol described previously, Immediately after getting rid of traces of genomic DNA by remedy with DNase I, an auto mated capillary gel electrophoresis was utilized to assess RNA high quality and quantity making use of a Bioanalyzer 2100 with RNA 6000 Nano Labchips.
On the dawn of a new paradigm in forest tree breeding, namely the
In the dawn of the new paradigm in forest tree breeding, namely the implementation of GS, a choice of factors that influences the accuracy of genomic estimated breeding values requires to become very carefully deemed, which includes the heritability in the traits, its genetic architecture, the extent of genotype by environment interaction, the genetic structure and also the productive size of the targeted population, the number of information inside the reference population, the number of markers and their linked price, as well as all round prediction and validation approach. The present examine supplies novel final results that need to be taken under consideration for your implementation of GS in maritime pine.
The drop inside the status variety from numerous hundred during the mass selected population, to 94 inside the second breeding population and 23 inside the elite more info here population of the new sub line structure with the breeding population is actually a favorable circumstance for its additional advancement within this species. Methods Genetic material, DNA extraction and genotyping assay The 2 mapping populations for which SNP based linkage maps have been merged in this research had been described in. G2 designates a 3 generation outbred pedigree, whereas F2 is usually a three generation inbred pedigree. Chancerel et al. constructed male and female linkage maps in the G2 population, along with a single linkage map for that F2 population, In addition, 194 trees in the base population of your maritime pine breeding program, referred to here since the 1st generation breeding or FGB population, have been applied for genetic diversity and LD examination.
Through the 1960s, grownup stands inside the Landes Motesanib forest had been explored and trees considered excellent when it comes to their stem volume and straightness had been recognized. These trees have been sampled across a broad range of distinct places covering the Aquitaine area, specifically along the Atlantic coast, and had been at least 50 m apart when current on the similar site. A phenotypic index was developed through the performances on the candidate trees and their 20 closest neighbors, to pick the base population. These trees were grafted and stored in clonal archives, Young needles from every single tree had been harvested and stored at 80 C right up until DNA extraction and genotyping, as described in, In total, 9,279 SNPs had been individually inspected with Genome Studio software, employing a GenCall score cutoff of 0. 15 to detect failed SNPs.
Loci for which two or 3 clusters had been recognized without the need of ambiguity had been viewed as to become polymorphic markers. SNP clusters were modified manually, to refine cluster positions, when necessary. SNPs and surrounding sequences were submitted to dbSNP, All round, 186 from the preliminary set of 194 trees presented genotyping information for 2,600 SNPs, Linkage map advancement We in contrast two different software program packages to make a composite map from three existing SNP primarily based linkage maps of maritime pine. LPmerge, that is out there as an R package at packages LPmerge, and MergeMap, which has been used in several barley mapping tasks, To assess the two algorithms, the root imply squared error for every marker was calculated by evaluating its place in the composite map with that on the personal linkage maps, as well as the regular RMSE throughout the markers within a linkage group was made use of to assess the goodness of match for your composite map.
